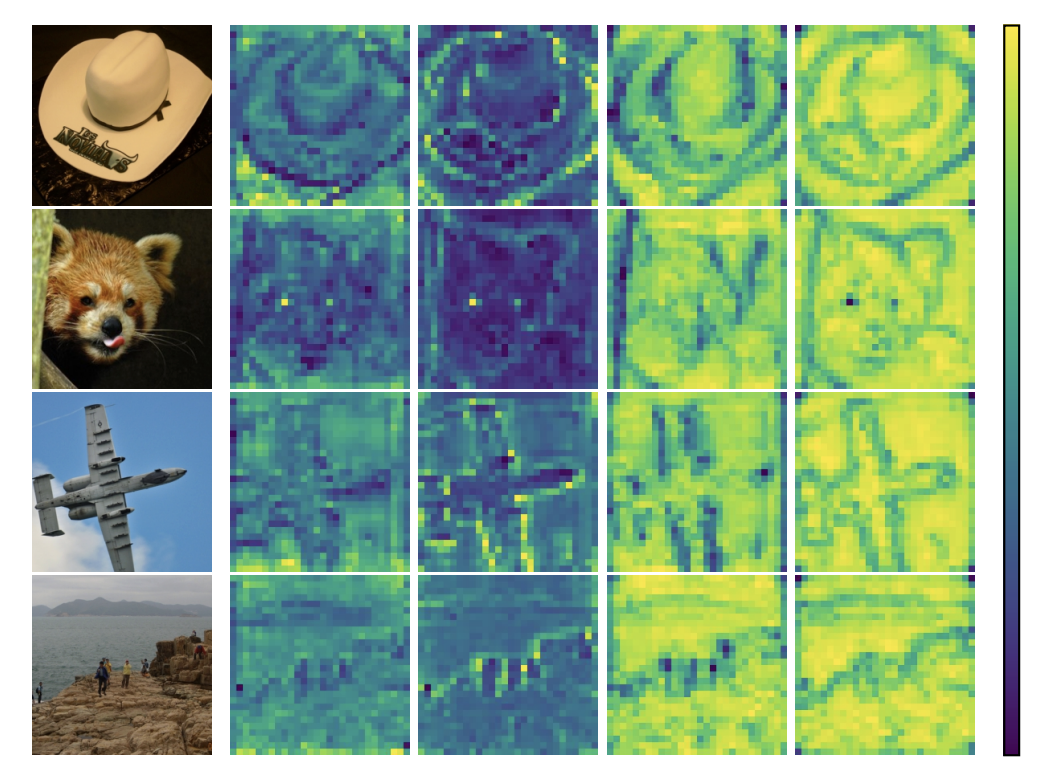
Involution是由Li等人于2021年提出的新型神经网络算子。该算子基于卷积(Convolution)设计而来,具有与卷积相反的基本特征,故得名,也常被称作内卷、合卷。本文统一称为 Involution。
本文记录笔者在本科毕业设计中对 Involution 的学习与实现过程,包括基础原理梳理、结构改造思路,以及并行化尝试与性能相关思考。
Involution is a neural network operator proposed by Li et al. in 2021. It is designed with characteristics opposite to standard convolution, and is often referred to as an inverse-style convolutional operator.
This post records my study and implementation notes from undergraduate thesis work, including core ideas, structural modifications, and early parallelization attempts for acceleration.
写在前面
本文涉及的技术论文作者目前因学术不端存在一定争议,但该篇论文本身当前没有被证实涉及争议内容。本文仅用作学习和记录,如果后续该篇论文或其技术被证明涉嫌学术问题,本文将永久归档或删除。
初步介绍
在机器学习中,卷积(Convolution)是非常经典的算法之一,特别是在计算机视觉(CV)领域。我们可以简单回顾一下二维卷积的操作,通过一个核(Kernel),对二维特征进行滑动提取特征。在深入了解Involution之前,我们需要理解传统卷积的一些特性。卷积操作通过共享的卷积核在空间位置上滑动,这种设计具有两个核心特征:空间不变性(spatial-agnostic)和通道特异性(channel-specific)。
空间不变性意味着同一个卷积核会应用到特征图的所有空间位置,无论位置在哪里,使用的权重都是相同的。这种设计在参数使用上效率较高,但也限制了模型对不同空间位置自适应建模的能力。通道特异性则体现在卷积会为每个输出通道学习独立的卷积核,这给卷积核带来冗余。
Involution算子则是基于与卷积相反的特性而设计的,它具有空间特异性(spatial-specific)和通道不变性(channel-agnostic)。这意味着Involution会为每个空间位置生成专属的核,这些核在通道维度上共享,并具有下面的优势:
- 自适应性增强:每个像素位置都有自己的处理核,能够更好地捕捉局部上下文信息;
- 参数效率提升:通道共享机制大幅减少了参数量,特别是在高通道数的深层网络中;
- 长距离依赖建模:通过增大核尺寸,Involution可以高效地建模远距离的空间关系,同时减少近距离信息丢失,平衡了远近规模的数据提取能力。
简单来说,这个算子在每个位置上生产自适应核,而通道中则通过分组确定哪些通道共用一个核心。这样就能每个元素使用不同核的同时,避免不同通道产生的冗余。接下来笔者会简单解释其原理。
原理概述
以二维数据为例,对于输入特征图
其中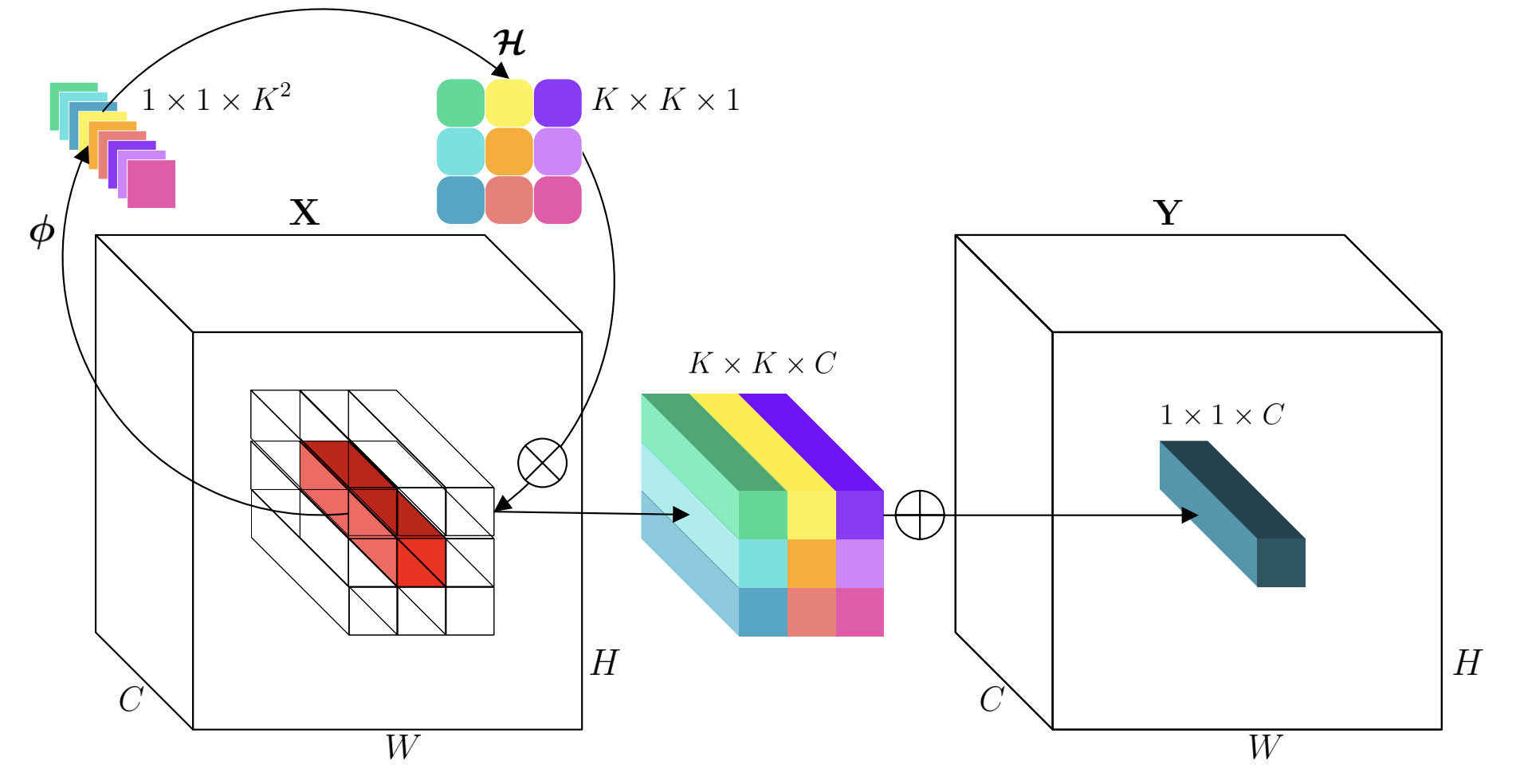 笔者这样理解:Involution基于单位置通道信息生成核,这样核参数本身就包含有通道特征,再通过通道共享,这样就能避免通道独立运算和输出时综合多通道的计算过程,从而减少计算量。
笔者这样理解:Involution基于单位置通道信息生成核,这样核参数本身就包含有通道特征,再通过通道共享,这样就能避免通道独立运算和输出时综合多通道的计算过程,从而减少计算量。
基本实现分析(程序)
上文提到了Involution的核生成函数,其使用两次线性变换来实现核权重的生成,那么具体怎么做呢?原论文中给出了实现的代码,尝试从代码的角度,分别观察两次变换的具体方法。
先来看看Pytorch风格的代码:
1 | |
在initialization的部分中,可以看到使用了两个Conv2d函数,切函数中规定的卷积核大小均为1。我们知道,
上述的两次变换是核生成的核心步骤,不过目前为止核心参数还没有体现为理论的三维形状,因此在forward
pass部分中,将数组进行了reshape。将输出的领域拼接成列,然后把列按照通道进行分组,重排维度以获得理想的核形状
当然,上面只是简单地分析基本的代码实现,了解到这一步,其实也能够根据项目的需求对算子进行调整了。
简单的项目应用
在笔者的项目中,使用Involution算子构建神经网络来预测患病概率,使用的是基于患者信息和临床数据表现作为特征的数据集(参考UCI心脏病数据集)。该数据集并非序列信息,但是可以通过卷积提取范围内属性的关联特征,而上文也提到,Involution算子的特性使其可以在大跨度数据中保留局部特征提取的能力,同时会有更少的计算量,因此也可以在非序列数据集中使用。不过,因为该数据集是一维的,所以不能直接使用原代码,需要对算子进行降维。
只要了解其基本的实现方法,降维的逻辑并不难,将原先的二维卷积操作修改为一维卷积即可。一维的involution算子逻辑可以参考下面的样例:
核生成的操作是基于通道进行的,与其数据本身的维度其实没有太多关联。对于高维数据集也可以推广,只是不能直接调用现有库了,需要自己实现高维的卷积操作。
当然,上面的样例中可以看到,笔者也修改了算子核生成步骤中的激活函数。原代码中使用的是ReLU,而笔者选择了能够对小负值更敏感的Swish。这里其实可以根据项目选择性调整,如果感觉involution算子所构建的网络表现不及预期,可以尝试调整激活函数。笔者的项目中,该项调整带来的提升比较小。
模型加速方案思考
笔者在完成另一项并行计算课程时的一个想法,既然involution的核生成是基于固定数据进行计算,那么它也可能进行并行加速。实际上,在论文作者的源代码中也已经给出了一个CUDA加速的版本。根据常规的并行化思路,可以将每一个位置的核生成操作并行。笔者尝试过分步并行方法,即两次卷积分别并行化矩阵运算,然后中间层进行等待。这种并行思路在CPU上运行时也能有较好的加速效果。不过这只是一次简单的尝试,毕竟大部分情况下也不会在CPU上进行模型训练。也许能有其它思路......
参考文档
Involution is a novel neural network operator proposed by Li et al. in 2021. This operator is designed based on Convolution, possessing fundamental characteristics opposite to those of convolution, hence its name. It is also often referred to as "内卷" (involution/internal scrolling) or "合卷" (combined convolution). This article will consistently refer to it as Involution.
This article records the author's learning and implementation process of Involution during their undergraduate thesis, including a review of the basic principles, ideas for structural modification, and attempts at parallelization along with performance-related considerations.
Involution is a neural network operator proposed by Li et al. in 2021. It is designed with characteristics opposite to standard convolution, and is often referred to as an inverse-style convolutional operator.
This post records my study and implementation notes from undergraduate thesis work, including core ideas, structural modifications, and early parallelization attempts for acceleration.
Foreword
The author of the technical paper involved in this article is currently under some controversy due to academic misconduct, but the paper itself has not been confirmed to be involved in the controversial content. This article is for learning and record-keeping purposes only. If the paper or its technology is later proven to be involved in academic issues, this article will be permanently archived or deleted.
Preliminary Introduction
In machine learning, Convolution is one of the most classic algorithms, especially in the field of Computer Vision (CV). We can briefly review the operation of 2D convolution, which extracts features by sliding a kernel over 2D features. Before delving into Involution, we need to understand some characteristics of traditional convolution. Convolutional operations slide a shared convolutional kernel across spatial locations. This design has two core features: spatial-agnosticism and channel-specificity.
Spatial-agnosticism means that the same convolutional kernel is applied to all spatial positions of the feature map; the weights used are identical regardless of the position. This design is highly efficient in terms of parameter usage but also limits the model's ability to adaptively model different spatial positions. Channel-specificity is reflected in convolution learning independent convolutional kernels for each output channel, which introduces redundancy in the kernels.
The Involution operator, on the other hand, is designed based on characteristics opposite to those of convolution: it possesses spatial-specificity and channel-agnosticism. This means that Involution generates a unique kernel for each spatial position, and these kernels are shared across the channel dimension, offering the following advantages:
- Enhanced adaptability: Each pixel location has its own processing kernel, allowing it to better capture local contextual information;
- Improved parameter efficiency: The channel-sharing mechanism significantly reduces the number of parameters, especially in deep networks with a high number of channels;
- Long-range dependency modeling: By increasing the kernel size, Involution can efficiently model long-range spatial relationships while reducing the loss of short-range information, balancing the ability to extract data at both close and distant scales.
Simply put, this operator produces adaptive kernels at each position, while in the channels, grouping determines which channels share a core. This allows each element to use a different kernel while avoiding redundancy produced by different channels. Next, I will briefly explain its principle.
Principle Overview
Taking 2D data as an example, for an input feature map \(\mathbf{X} \in \mathbb{R}^{H \times W \times C}\), the output of Involution at position \((i,j)\) can be expressed as:
\[ Y_{i,j}= \sum_{(u,v)\in\mathcal{N}(i,j)} \mathcal{H}_{i,j,u,v} \cdot \mathbf{X}_{u,v} \]
where \(\mathcal{N}(i,j)\) is the
neighborhood centered at \((i,j)\), and
\(\mathcal{H}_{i,j} \in \mathbb{R}^{K \times K
\times C}\) is the involution kernel for that position. Its
kernel mapping function is defined as: \[
\phi = \mathbb{R}^C \mapsto \mathbb{R}^{K\times K\times G}
\] This expression means that the kernel mapping, with channel as
the dimension, takes a three-dimensional form of kernel size multiplied
by the number of groups. It can be understood that the kernel mapping
operation is performed based on the channel vector corresponding to the
element's position. The following function is its specific expression:
\[
\mathcal{H}_{i,j} =
\phi(\mathbf{X}_{i,j})=W_1\sigma(W_0\mathbf{X}_{i,j})
\] Here, \(W_0\) and \(W_1\) are two linear transformations, and
\(\sigma\) refers to the activation
function. After obtaining the kernel parameters through the above
mapping, they are then applied to the original data to complete the
kernel operation. Below is the schematic diagram used in the original
paper:  My understanding is this: Involution generates
kernels based on single-position channel information. This way, the
kernel parameters themselves contain channel features. Then, through
channel sharing, it avoids independent channel operations and the
multi-channel integration process during output, thereby reducing
computational complexity.
My understanding is this: Involution generates
kernels based on single-position channel information. This way, the
kernel parameters themselves contain channel features. Then, through
channel sharing, it avoids independent channel operations and the
multi-channel integration process during output, thereby reducing
computational complexity.
Basic Implementation Analysis (Code)
The Involution kernel generation function mentioned above uses two linear transformations to generate kernel weights. How exactly is this done? The original paper provides the implementation code; let's observe the specific methods of these two transformations from a code perspective.
First, let's look at the PyTorch-style code:
1 |
|
In the initialization section, we can see that two Conv2d functions are used, and the kernel size specified in these functions is 1. We know that a \(1\times 1\) convolutional kernel operation only involves the channel dimension of the data and no other dimensional information. In other words, these functions perform a linear combination for the channels corresponding to each position. Combining this with the formulas discussed earlier, these two 2D convolution functions with kernel size 1 are actually the linear transformations \(W_0\) and \(W_1\) mentioned above, which can be written in the form: \[ \begin{align*} &W_0:\mathbb{R}^C \mapsto \mathbb{R}^{C//r}\\ &W_1:\mathbb{R}^{C//r} \mapsto \mathbb{R}^{K\times K\times G}\\ \end{align*} \] The first transformation \(W_0\) performs dimensionality reduction, used to decrease the number of calculated kernel parameters; the second transformation \(W_1\) implements mapping, transforming the compressed features into the required number of weight values.
The two transformations above are the core steps of kernel generation. However, so far, the core parameters have not yet taken on the theoretical three-dimensional shape. Therefore, in the forward pass section, the array is reshaped. The output neighborhoods are concatenated into columns, and then these columns are grouped by channel, and dimensions are reordered to obtain the desired kernel shape \((B,G,K\times K,H,W)\). Here, \(H\) and \(W\) refer to the output position information. Finally, it is applied to the full channel dimension to obtain the final feature map.
Of course, the above is just a simple analysis of the basic code implementation. Understanding this step already allows for adjustments to the operator based on project requirements.
Simple Project Application
In my project, I used the Involution operator to build a neural network for predicting disease probability, utilizing a dataset based on patient information and clinical data as features (refer to the UCI Heart Disease Dataset). This dataset is not sequential information, but convolutional operations can extract associated features within attribute ranges. As mentioned earlier, the characteristics of the Involution operator allow it to retain local feature extraction capabilities in large-span data with less computational overhead, making it suitable for non-sequential datasets. However, since this dataset is one-dimensional, the original code cannot be used directly, and the operator needs to be downscaled.
As long as one understands its basic implementation method, the logic
for dimensionality reduction is not difficult: simply modify the
original 2D convolution operation to a 1D convolution. The logic for a
1D involution operator can be seen in the example below:
The kernel generation operation is performed based on channels and is not directly related to the intrinsic dimensionality of the data itself. This can also be generalized to higher-dimensional datasets, although existing libraries cannot be called directly, and high-dimensional convolutional operations would need to be implemented manually.
Of course, as seen in the example above, I also modified the activation function in the operator's kernel generation step. The original code used ReLU, while I chose Swish, which is more sensitive to small negative values. This can be selectively adjusted based on the project; if the network built with the involution operator performs below expectations, one can try adjusting the activation function. In my project, this adjustment brought a relatively small improvement.
Thoughts on Model Acceleration Solutions
An idea I had while completing another parallel computing course was that since Involution's kernel generation is based on fixed data, it might also be amenable to parallel acceleration. In fact, the paper's authors have already provided a CUDA-accelerated version in their source code. Following conventional parallelization ideas, the kernel generation operation for each position can be parallelized. I experimented with a step-by-step parallelization method, where the two convolutional operations are parallelized as matrix computations separately, with an intermediate layer waiting. This parallel approach showed good acceleration even when running on a CPU. However, this was just a simple attempt, as model training usually isn't performed on a CPU. Perhaps there are other ideas...